RESEARCH
I am the Director of Computational Genomics at AbbVie. As a scientist , I am intrigued by the application of high throughput omics technologies to accelerate drug discovery and advance precision medicine. Before this, I was Director of Bioinformatics at Lurie Children's Hospital of Chicago and Assistant Professor of Pathology at Northwestern University. Here, I led a team of bioinformaticians and scientists to design, develop and implement cloud-based computational infrastructure and containerized bioinformatics software for clinical diagnostic services for germline and somatic next generation sequencing (NGS) testing. I worked on informatics solutions and research questions for translational ‘omics’ and biomedical research to promote precision medicine goals. At the Broad Institute, I worked with large non-coding RNAs and developed a software to mine end RNA-sequencing data. During my PhD, I studied the evolution of miRNA regulation involved in developmental gene regulatory networks in echinoderms, especially, sea urchin and sea stars.
My PhD thesis: miRNA regulation in development
My Google Scholar profile can be found HERE.
Check out this article about my work on GenomeWeb.
Software
Selected Publications
My Pubmed bibliography can be found HERE.
Ritterhouse LL, Parilla M, Zhen CJ, Wurst MN, Puranik R, Henderson CM, Joudeh NZ, Hartley MJ, Haridas R, Wanjari P, Furtado LV, Kadri S, Segal JP. Clinical Validation and Implementation of a Measurable Residual Disease Assay for NPM1 in Acute Myeloid Leukemia by Error-Corrected Next-Generation Sequencing. Mol Diagn Ther. 2019 Oct 31.
Balagopal V, Hantel A, Kadri S, Steinhardt G, Zhen CJ, Kang W, Wanjari P, Ritterhouse LL, Stock W, Segal JP. Measurable residual disease monitoring for patients with acute myeloid leukemia following hematopoietic cell transplantation using error corrected hybrid capture next generation sequencing. PLoS One. 2019 Oct 28;14(10):e0224097.
Choudhury NJ, Eghtesad M, Kadri S, Cursio J, Ritterhouse L, Segal J, Husain A, Patel JD. Fewer actionable mutations but higher tumor mutational burden characterizes NSCLC in black patients at an urban academic medical center. Oncotarget. 2019 Oct 8;10(56):5817-5823.
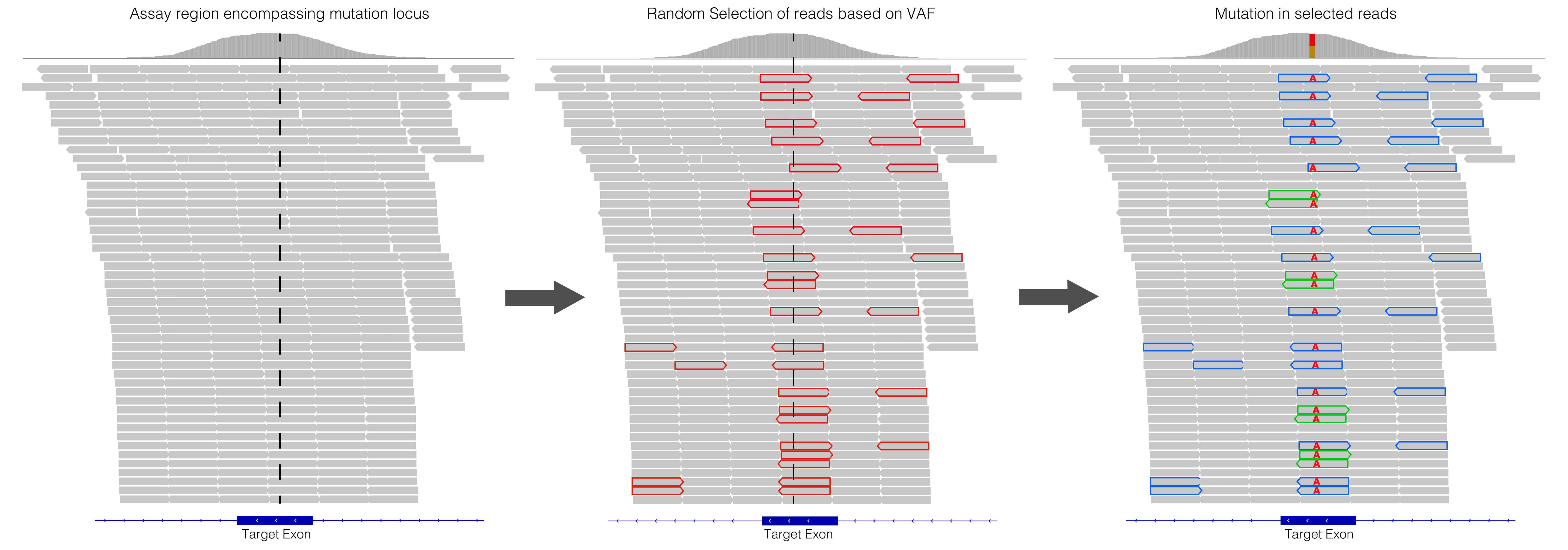
Patil SA, Mujacic I, Ritterhouse LL, Segal JP, Kadri S.insiM: in silico Mutator Software for Bioinformatics Pipeline Validation of Clinical Next-Generation Sequencing Assays. J Mol Diagn. 2019 Jan;21(1):19-26.
Kang W*, Kadri S*, Puranik R, Wurst MN, Patil SA, Mujacic I, Benhamed S, Niu N, Zhen CJ, Ameti B, Long BC, Galbo F, Montes D, Iracheta C, Gamboa VL, Lopez D, Yourshaw M, Lawrence CA, Aisner DL, Fitzpatrick C, McNerney ME, Wang YL, Andrade J, Volchenboum SL, Furtado LV, Ritterhouse LL, Segal JP.System for Informatics in the Molecular Pathology Laboratory: An Open-Source End-to-End Solution for Next-Generation Sequencing Clinical Data Management.J Mol Diagn. 2018 Jul;20(4):522-532.
Kadri S, Lee J, Fitzpatrick C, Galanina N, Sukhanova M, Venkataraman G, Sharma S, Long B, Petras K, Theissen M, Ming M, Kobzev Y, Kang W, Guo A, Wang W, Niu N, Weiner H, Thirman M, Stock W, Smith SM, Nabhan C, Segal JP, Lu P, Wang YL. Clonal evolution underlying leukemia progression and Richter transformation in patients with ibrutinib-relapsed CLL.. Blood Adv. 2017 May 2;1(12):715-727.
Kadri S., Long BC, Mujacic I, Zhen CJ, Wurst MN, Sharma S, McDonald N., Niu N, Benhamed S, Tuteja J, Seiwert T, White KP, McNerney ME, Fitzpatrick C, Wang YL, Furtado LV, Segal JP . UCM-OncoPlus: Clinical Validation of a Comprehensive Genomic Oncology Panel Via Cross-Platform Comparison with Established Next Generation Sequencing Assays J Mol Diagn. 2017 Jan;19(1):43-56. doi: 10.1016/j.jmoldx.2016.07.012. Epub 2016 Nov 9.
Chen J, Shishkin AA, Zhu X,Kadri S., Maza I, Guttman M, Hanna JH, Regev A, Garber M. Evolutionary analysis across mammals reveals distinct classes of long non-coding RNAs. Genome Biol. 2016 Feb 2;17(1):19. doi: 10.1186/s13059-016-0880-9.
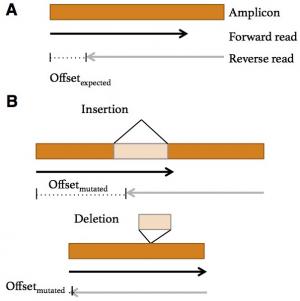
Kadri S., Zhen CJ., Wurst M.N., Long B.C., Jiang Z., Wang Y.L., Furtado L.V., Segal J.P. Amplicon Indel Hunter: A Novel Bioinformatics Tool to Detect Large Somatic Insertion/Deletion Mutations in Amplicon-Based NGS Data. J Mol Diagn. 2015 Nov;17(6):635-43. doi: 10.1016/j.jmoldx.2015.06.005. Epub 2015 Aug 28.
Suvà M., Rheinbay E., Gillespie S., Patel A., Wakimoto H., Rabkin S., Chi A.,Cahill D., Nahed B., Curry W., Martuza R., Rivera M., Riggi N., Rosetti N., Kasif S., Beik S., Kadri S., Tirosh I., Wortman Ivo, Shalek A., Rozenblatt-Rosen O., Regev A., Loius D., Bernstein B. 2014 Reconstructing and reprogramming the tumor propagating potential of glioblastoma stem-like cells. Cell. 2014 Apr 24;157(3):580-94
Engreitz J., Pandya-Jones A., McDonel P., Shishkin A., Sirokman K., Surka C., Kadri S., Xing J., Goren A., Lander E., Plath K., and Guttman M. 2013 The Xist lncRNA Exploits Three-Dimensional Genome Architecture to Spread Across the X Chromosome. Science Vol. 341 no. 6147 doi: 10.1126/science.1237973
Kadri S., Hinman V., and Benos P. 2011 RNA deep sequencing reveals differential microRNA expression during development of sea urchin and sea star. PLoS ONE 6(12): e29217. doi:10.1371/journal.pone.0029217
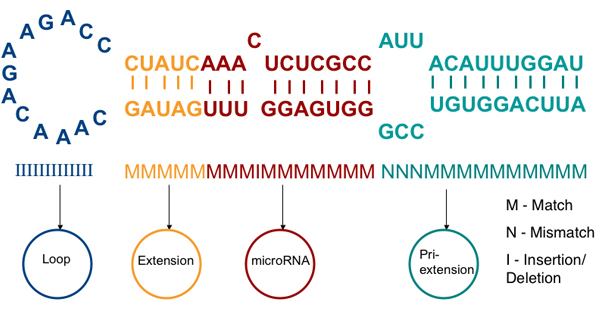
Kadri S., Hinman V., and Benos P. 2009 HHMMiR: efficient de novo prediction of microRNAs using hierarchical hidden Markov models. BMC Bioinformatics 10, no. 1: S35. doi:10.1186/1471-2105-10-S1-S35
PhD research
I have developed a probabilistic model for microRNA precursor prediction, called HHMMiR based on hierarchical hidden Markov models (HHMMs), without requirement of conservation of sequence between closely related species. It can be downloaded here .
We studied small RNA libraries in sea uchin and sea star embryonic samples to study microRNA populations.
Whole mount in situ hybridization to study spatial and temporal microRNAs expression patterns, and knocked down critical genes involved in the microRNAs biogenesis pathway to knock down microRNA function in early sea urchin embryos and studied effects on embryonic development.
My PhD thesis: miRNA regulation in development